
The REPLI-g Advanced DNA Single Cell Kit features a simplified protocol of just 3 steps with only 20 minutes hands-on time for reaction setup and 2 hours for reaction time. The typical yield from a single cell is 25–35 µg of amplified DNA – sufficient for downstream applications involving qPCR and next-generation sequencing. The REPLI-g Advanced DNA Single Cell protocol saves at least 1 hour of time versus the first-generation REPLI-g Single Cell Kit (cat. nos. 150343 and 150345).
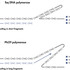

[A] Upon encountering secondary DNA structures, Taq polymerase may pause synthesis, slip, or dissociate from the template. This can result in inaccurate DNA amplification, incomplete loci coverage, and short fragment sizes. [B] REPLI-g Kits utilize Phi 29 polymerase, which displaces secondary structures enabling accurate and highly uniform amplification of the entire genome.
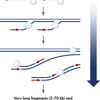
Primers (arrows) anneal to the template DNA and are extended at 30°C by Phi 29 polymerase, which moves along the DNA template strand, displacing the complementary strand, while becoming a template itself for replication. In contrast to PCR amplification, MDA does not require different temperatures and ends in very long fragments with low mutation rates.
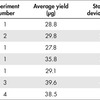
REPLI-g Advanced DNA Single Cell Kits were tested in multiple experiments by 2 operators, using 4 different lots of reagents, with 4 different types of eukaryotic cells. Single cells were isolated with the QIAGEN QIAscout.
For each experiment, 6000 cells were loaded onto the QIAscout array and left overnight in a standard cell incubator. For suspension cells, the QIAscout array was first treated with Cell-Tak to facilitate adherence of the cells. On the day of cell collection, QIAscout was installed on a 10x objective of an inverted microscope (Axio Vert.A1, Zeiss [Carl Zeiss AG]) and the array was screened using brightfield microscopy. Microrafts containing single cells were dislodged and transferred to a secondary vessel containing 4 µl REPLI-g Advanced sc Storage Buffer prior to starting the REPLI-g Advanced DNA Single Cell protocol for single cells.
Data presented is from 7 separate experiments (4 experiments with Jurkat cells and 3 experiments with additional cell lines) and 4 separate lots of reagents (A–D). Final DNA yield was quantified using PicoGreen (Thermo Fisher Scientific). Other quantification methods that are not specific for dsDNA might overestimate the yield.
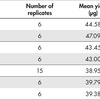
REPLI-g Advanced DNA Single Cell Kits were used to amplify 1 ng human genomic DNA. Two different operators (DU and AJ) and different lots of Reaction Buffer (A–E) were used in 7 separate experiments to confirm protocol and reagent reproducibility.
Final DNA yield was quantified using PicoGreen (Thermo Fisher Scientific). Other quantification methods that are not specific for dsDNA might overestimate the yield because REPLI-g Advanced DNA Single Cell amplification products might contain unused reaction primers and nucleotides.

REPLI-g–amplified DNA from isolated single Jurkat cells using the first generation REPLI-g Single Cell Kit (i.e., Original REPLI-g Single Cell Kit) and the new second generation REPLI-g Advanced DNA Single Cell Kit was analyzed by qPCR using both single (DOLK, C4:00, C4:30, C4:40, C4:60, C4:80 and 3q) and multi-copy (Multi) loci. The multi-copy gene provides information regarding the overall success of the WGA (whole-genome amplification) while the single-copy genes give an estimation about uniformity of WGA. A major increase in the C T value for single copy genes is indicative of potential allelic dropout.
The results show improved robustness and reproducibility with the REPLI-g Advanced DNA Single Cell Kit. The simple 2 hour amplification reaction is adequate for obtaining sufficient amplified DNA for qPCR applications, while the improved reaction conditions limit allelic dropout.

REPLI-g–amplified DNA from isolated single LoVo cells using the first generation REPLI-g Single Cell Kit (i.e., Original REPLI-g Single Cell Kit) and the new second generation REPLI-g Advanced DNA Single Cell Kit was analyzed by qPCR using both single (DOLK, C4:00 and 3q) and multi-copy (Multi) loci. The multi-copy gene provides information regarding the overall success of the WGA (whole-genome amplification) while the single-copy genes give an estimation about uniformity of WGA. A major increase in the C T value for single copy genes is indicative of potential allelic dropout.
The results show improved robustness and reproducibility with the REPLI-g Advanced DNA Single Cell Kit. The simple 2 hour amplification reaction is adequate for obtaining sufficient amplified DNA for qPCR applications, while the improved reaction conditions limit allelic dropout.

REPLI-g–amplified DNA from isolated single MCF7 cells using the first generation REPLI-g Single Cell Kit (i.e., Original REPLI-g Single Cell Kit) and the new second generation REPLI-g Advanced DNA Single Cell Kit was analyzed by qPCR using both single (DOLK, C4:00 and 3q) and multi-copy (Multi) loci. The multi-copy gene provides information regarding the overall success of the WGA (whole-genome amplification) while the single-copy genes give an estimation about uniformity of WGA. A major increase in the C T value for single copy genes is indicative of potential allelic dropout.
The results show improved robustness and reproducibility with the REPLI-g Advanced DNA Single Cell Kit. The simple 2 hour amplification reaction is adequate for obtaining sufficient amplified DNA for qPCR applications, while the improved reaction conditions limit allelic dropout.
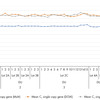
REPLI-g Advanced DNA Single Cell Kits were used to amplify 1 ng human genomic DNA. Two operators (DU and AJ), different DNA Polymerase lots (2–5) and different lots of Reaction Buffer (A–C) were used to confirm protocol and reagent reproducibility. REPLI-g–amplified gDNA was analyzed by qPCR using both single (DOLK and C4:00) and multi-copy (Multi) loci. The multi-copy gene provides information regarding the overall success of the WGA (whole-genome amplification) while the single-copy genes give an estimation about uniformity of WGA. A major increase in the C T value for single copy genes, when compared between WGA reactions, is indicative of potential allelic dropout.
The results show REPLI-g Advanced DNA Single Cell Kits have a high degree of reproducibility regardless of the operator or lots of reagents used. The simple 2 hour amplification reaction is adequate for obtaining sufficient amplified gDNA for qPCR applications, while the improved reaction conditions limit allelic dropout.
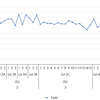
REPLI-g Advanced DNA Single Cell Kits were used to amplify 1 ng human genomic DNA. Two operators (DU and AJ), different DNA Polymerase lots (2–5) and different lots of Reaction Buffer (A–C) were used to confirm protocol and reagent reproducibility. Yield of WGA was measured using PicoGreen (Thermo Fisher Scientific) reagent in conjunction with a fluorometer. PicoGreen reagent displays enhanced binding to double-stranded DNA and is therefore suitable to quantify the double-stranded DNA product. Other quantification methods that are not specific for dsDNA might overestimate the yield because REPLI-g Advanced DNA Single Cell amplification products might contain unused reaction primers and nucleotides. The average yield across all experiments is 41 µg, which is an amplification of ~40,000-fold.
For PicoGreen measurement, the protocol described in the Appendix of the REPLI-g Advanced DNA Single Cell Kit Handbook was used.
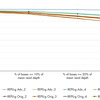
Single Jurkat cells were isolated using the QIAscout and transferred to a secondary vessel containing 4 µl REPLI-g Advanced sc Storage Buffer or 4 µl PBS. REPLI-g Advanced sc Storage Buffer is a feature of the new kit. It preserves cell integrity prior to cell lysis and helps reduce DNA damage.
Single cells in 4 µl of REPLI-g Advanced sc Storage Buffer were processed with the REPLI-g Advanced DNA Single Cell Kit (REPLI-g Adv) while single cells in 4 µl of PBS were processed with the REPLI-g Single Cell Kit (REPLI-g Orig). Amplification yields were quantified using PicoGreen (Thermo Fisher Scientific). 40 ng of amplified DNA from single cells, as well as unamplified gDNA from bulk Jurkat cells, was used to build targeted NGS libraries using the QIAseq Targeted DNA Human Comprehensive Cancer Panel (DHS-3501Z) covering 275 genes (837 kb). Sequencing was performed on the MiSeq (Illumina).
To evaluate uniformity, calculations were made for how many target bases reach 5–30% (at minimum) of the mean read depth. The new REPLI-g Advanced DNA Single Cell Kit resulted in higher uniformity than placing cells in PBS and using the original REPLI-g Single Cell Kit.
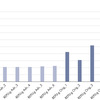
Single allelic dropout (ADO) for REPLI-g–amplified DNA from single cells was assessed using libraries generated using the QIAseq Targeted DNA Human Comprehensive Cancer Panel and sequenced using a MiSeq (Illumina). A comparison between 6 single Jurkat cells using the REPLI-g Advanced DNA Single Cell kit and 6 single Jurkat cells using the original REPLI-g Single Cell Kit is shown. Unamplified gDNA from bulk Jurkat cells was used as reference. NGS results were analyzed using the GeneGlobe Data Analysis Center (www.qiagen.com/shop/genes-and-pathways/data-analysis-center-overview-page/).
Each library is based on a single cell after WGA, with 40 ng amplified DNA used as starting material, and a mean read depth of 127. To compare the ADO rates for both kits, we analyzed the single ADO for SNPs that are heterozygous (frequency of SNP allele: 25 – 75 %) in the unamplified bulk DNA. Calculation of ADO rate is: (# of heterozygous loci in gDNA – # of heterozygous loci in single cell after WGA)/(# of heterozygous loci in gDNA).
The REPLI-g Advanced DNA Single Cell Kit shows a lower rate of ADO and improved inter-sample consistency.

Single Jurkat cells captured using the QIAscout were used in MALBAC Single Cell WGA Kit (Yikon Genomics), PicoPLEX WGA Kit (Rubicon Genomics, Inc.) and REPLI-g Advanced DNA Single Cell Kit (QIAGEN) amplifications according to the protocol provided with each kit. Amplified DNA was analyzed using qPCR for single (DOLK, C4:00 and 3q) and multi-copy (Multi) loci. The multi-copy gene provides information regarding the overall success of the WGA (whole-genome amplification) while the single-copy genes give an estimation about uniformity of WGA. A major increase in the C T value for single copy genes is indicative of potential allelic dropout.
The results show improved robustness and reproducibility with the REPLI-g Advanced DNA Single Cell Kits versus other commercially-available kits. The simple 2 hour amplification reaction is adequate for obtaining sufficient amplified gDNA for qPCR applications, while the improved reaction conditions limit allelic dropout.
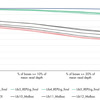
Single Jurkat cells were collected using the QIAscout. Isolated single cells on QIAscout microrafts were transferred to a secondary vessel containing 4 μl REPLI-g Advanced sc Storage Buffer (REPLI-g Advanced DNA Single Cell Kit), 5 µl Extraction Cocktail (PicoPLEX WGA Kit [Rubicon Genomics]) or 5 µl Lysis Reaction Mix (MALBAC Single Cell WGA Kit [Yikon Genomics]).
40 ng of amplified DNA, as well as unamplified gDNA (isolated from bulk Jurkat cells), was used to build targeted NGS libraries using the QIAseq Targeted DNA Human Comprehensive Cancer Panel (DHS-3501Z). This panel has full exon coverage of 275 genes with coverage of 837 kb. Sequencing was performed on the MiSeq (Illumina).
To evaluate uniformity, calculations were made for how many target bases reach 5–30% (at minimum) of the mean read depth. The combination of the REPLI-g Advanced sc Storage Buffer and the improved chemistry of the REPLI-g Advanced DNA Single Cell Kit results in higher uniformity compared to alternative methods.